Using BEAGLE with BEAST
BEAGLE is a high-performance library that can perform the core calculations at the heart of most Bayesian and Maximum Likelihood phylogenetics package. It can make use of highly-parallel processors such as those in graphics processing units (GPUs) found in many PCs.
The current version of BEAGLE will only work with BEAST v1.6 or later
Downloading and installing BEAGLE
BEAGLE is an independent project and its website is https://github.com/beagle-dev/beagle-lib, where binary installers and installation instructions for different operating systems are available.
Testing the BEAGLE installation
To test the installation, run BEAST and when the options dialog box appears, select “Use BEAGLE library” and “Show list of available BEAGLE resources”:
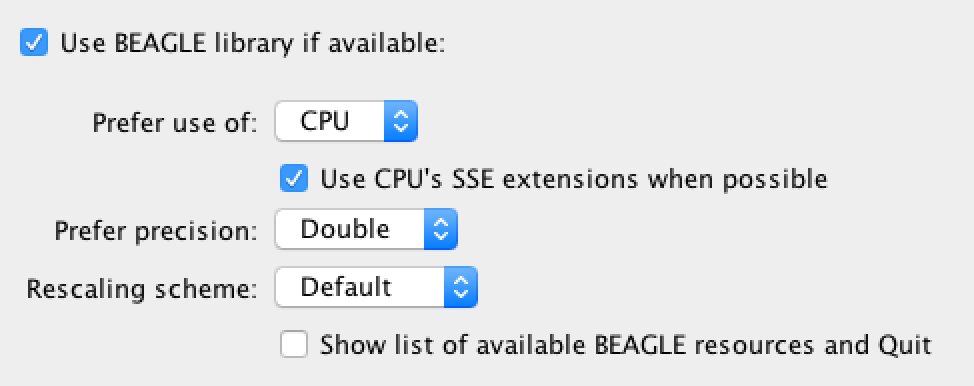
You don’t need to specify a BEAST input file as the information about BEAGLE will be displayed and then BEAST will stop. For the commandline version of BEAST, type:
beast -beagle_info
When run, it should show a list of available resources that BEAGLE has detected:
BEAGLE resources available:
0 : CPU
Flags: PRECISION_SINGLE PRECISION_DOUBLE ...
1 : Intel(R) HD Graphics 530
Global memory (MB): 1536
Clock speed (Ghz): 1.05
Number of compute units: 24
Flags: PRECISION_SINGLE COMPUTATION_SYNCH ...
2 : Tesla K40c
Global memory (MB): 11520
Clock speed (Ghz): 0.74
Number of cores: 2880
Flags: PRECISION_SINGLE PRECISION_DOUBLE ...
This shows the CPU (which should always be available), an Intel integrated graphics processor (this is the graphics chip that is used to run the computer display) and one NVidia Tesla GPU card specifically designed for scientific computing.
Running BEAST with BEAGLE
You are now ready to run BEAST using BEAGLE. Just run BEAST again and select “Use BEAGLE library” but not the “Show list of available BEAGLE resources” option. When running, as part of the initial pre-amble text, BEAGLE will say what resource it is using:
Using BEAGLE TreeLikelihood
Branch rate model used: strictClockBranchRates
Using BEAGLE resource 0: CPU
with instance flags: PRECISION_DOUBLE COMPUTATION_SYNCH EIGEN_REAL SCALING_MANUAL SCALERS_RAW VECTOR_SSE THREADING_NONE PROCESSOR_CPU FRAMEWORK_CPU
Ignoring ambiguities in tree likelihood.
With 191 unique site patterns.
Using rescaling scheme : dynamic (rescaling every 100 evaluations, delay rescaling until first overflow)
This shows that BEAGLE will be using the primary CPU and gives some details about the options it will be using. The default options will be optimal for most machines but see this page for information about optimizing BEAST computational performance.
Running BEAST/BEAGLE on command line
If you are running BEAST on Linux or wish to have more control of the BEAGLE performance options on Mac OS X or Windows you will need to run BEAST from the command line.
Running BEAST from the command line allows you to select which resource needs to be used for your analysis by using a command-line option. For example, in order to ensure that resource 1 (see above) will be used when running BEAST, you can use the following command:
beast -beagle_gpu -beagle_order 1 data.xml
The full list of command-line BEAGLE options can be listed using beast -help and are listed here:
-beagle_off Dont use the BEAGLE library
-beagle Use BEAGLE library if available (default on)
-beagle_info BEAGLE: show information on available resources
-beagle_order BEAGLE: set order of resource use
-beagle_instances BEAGLE: divide site patterns amongst instances
-beagle_CPU BEAGLE: use CPU instance
-beagle_GPU BEAGLE: use GPU instance if available
-beagle_SSE BEAGLE: use SSE extensions if available
-beagle_SSE_off BEAGLE: turn off use of SSE extensions
-beagle_cuda BEAGLE: use CUDA parallization if available
-beagle_opencl BEAGLE: use OpenCL parallization if available
-beagle_single BEAGLE: use single precision if available
-beagle_double BEAGLE: use double precision if available
-beagle_async BEAGLE: use asynchronous kernels if available
-beagle_scaling BEAGLE: specify scaling scheme to use
-beagle_rescale BEAGLE: frequency of rescaling (dynamic scaling only)
Frequently Asked Questions
-
When attempting to run BEAST with BEAGLE, I get the following error message and BEAST/BEAGLE crashes:
# A fatal error has been detected by the Java Runtime Environment: # # EXCEPTION_ACCESS_VIOLATION (0xc0000005) at pc=0x476bcb8d, pid=2548, tid=0x00000244 # # JRE version: Java(TM) SE Runtime Environment (8.0_144-b01) (build 1.8.0_144-b01) # Java VM: Java HotSpot(TM) Client VM (25.144-b01 mixed mode windows-x86 ) # Problematic frame: # C [hmsbeagle-cpu-sse32-21.DLL+0xcb8d] # # Failed to write core dump. Minidumps are not enabled by default on client versions of Windows # # If you would like to submit a bug report, please visit: # http://bugreport.java.com/bugreport/crash.jsp # The crash happened outside the Java Virtual Machine in native code. # See problematic frame for where to report the bug.This isn’t so much a BEAGLE error as it is a compatibility problem with the operating system and the Java installation. Notice how the error message mentions a mixed mode installation, which usually indicates that a 32-bit Java version was installed on a 64-bit Windows operating system. Upgrading the 32-bit Java installation to a 64-bit version usually fixes this problem.
-
Upon building the BEAGLE library from source, I get the following error message when running the
configurescript:configure: error: cannot find JDK header filesThis can be fixed by correctly setting the
JAVA_HOMEvariable. For example, on a MacBook running Java version 8 Update 121, this can be done as follows:export JAVA_HOME=/Library/Java/JavaVirtualMachines/jdk1.8.0_121.jdk/Contents/Home/ -
When running BEAST using the installation packages (i.e. BEAST v1.10.0.app on Mac OS and BEAST v1.10.0.exe on Windows), the BEAGLE installation cannot be found:
Failed to load BEAGLE library: no hmsbeagle-jni in java.library.pathPossibly, the issue here is that you’re using the installation packages for BEAST but have installed BEAGLE from its source files (i.e. not using BEAGLE’s installation package). At the moment, it’s indeed not readily possible to combine these two approaches and we suggest to either use the installation packages for both BEAST and BEAGLE or to build both from source.
Citing BEAGLE
A manuscript describes the BEAGLE API and library: http://sysbio.oxfordjournals.org/content/61/1/170
The paper describing the algorithms used for calculating likelihoods of sequences on trees using many core devices like graphics processing units (GPUs) is available from: https://academic.oup.com/bioinformatics/article/25/11/1370/332982